Team I Webserver Group: Difference between revisions
No edit summary |
|||
| (38 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
==Introduction== | ==Introduction== | ||
===Background=== | ===Background=== | ||
The | The objectives of our BIOL 7210: Computational Genomics teams were to, given unassembled genome sequence data from the Weiss Lab at the Emory University School of Medicine, proceed through five distinct stages of analysis and interpretation of that data: genome assembly, gene prediction, functional annotation, comparative genomics, and production of a predictive webserver. At the last stage, our goal was to create a predictive webserver that performed the functionalities of some, if not all, of the work from previous groups. | ||
===Goals=== | |||
Our goals for a predictive webserver were as follows: | |||
*Assemble input reads | *Assemble input reads | ||
*Analyze assemblies | *Analyze assemblies | ||
*Visualize results | *Visualize results in user-friendly format | ||
*Implement a way for results to be downloaded | *Implement a way for results to be downloaded | ||
===KAREN=== | |||
'''K'''lebsiella '''A'''ntibiotics '''RE'''sistance Predicitio'''N''' (KAREN) is a culmination of these objectives and is able to perform the following analyses given an input of raw sequence reads: | |||
* De novo assembly | |||
* Species identification | |||
* Strain identification | |||
* Average Nucleotide Identity | |||
* Computational phenotyping | |||
* Visualization of results | |||
===Technologies Used=== | ===Technologies Used=== | ||
[[File:laravel.png]] | |||
For the creation and development of this webserver, we used PHP framework for server-side programming. PHP provides a strong frameworks to support MySQL and Apache Server. Also, PHP provides the feasibility of the development of Model-View-Controller (MVC) framework, which provides a more simple user-interface. There are many MVC frameworks available, among which we used Laravel. Laravel was used because of its extensive documentation and a large community of developers. It also currently is one of the most widely used PHP frameworks. For more information, please visit their [https://laravel.com/ website] | |||
This webserver is built on PHP v7.0.0 and Laravel v5.5. | |||
==Functionalities== | ==Functionalities== | ||
=== | ===De novo Genome Assembly using SKESA=== | ||
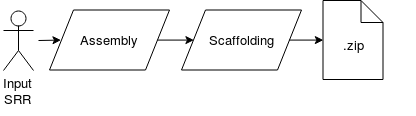
We used SKESA for de novo genome assembly. The input to the assembler was raw reads (forward & reverse) retrieved using SRA accession numbers. The output contigs then were scaffolded using a tool called SSPACE. For more information on the assembly pipeline, refer to [http://compgenomics2018.biosci.gatech.edu/Team_I_Genome_Assembly_Group Genome Assembly team]. SKESA is currently unpublished. | |||
[[File:Assembly_flowchart.png]] | |||
===Species & Strain Typing by StrainSeeker=== | ===Species & Strain Typing by StrainSeeker=== | ||
Strainseeker is a tool which lets you rapidly and accurately make an assessment of the species and strain of a bacterial assembly. StrainSeeker has a pre-built database that is uses for species identification and works on paired-end reads to identify strain type. It has the ability to identify novel strains and is therefore a useful tool for further assessment of a sample of unknown origin. | |||
<code>perl builder.pl -n refseq_guide_tree.nwk -d strain_fasta_directory -w 32 -o my_database</code> | <code>perl builder.pl -n refseq_guide_tree.nwk -d strain_fasta_directory -w 32 -o my_database</code> | ||
| Line 39: | Line 48: | ||
<code>perl seeker.pl -i sample_file.fastq -d ss_db_w32 -o sample_result.txt</code> | <code>perl seeker.pl -i sample_file.fastq -d ss_db_w32 -o sample_result.txt</code> | ||
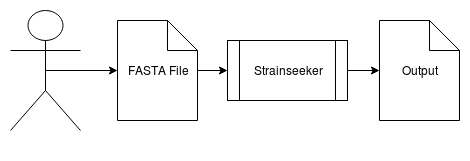
The pipeline works by taking a FASTA file as an input, which will then be processed by StrainSeeker. The output will be parsed by one of the scripts in the pipeline and visualization with tables containing information is generated. | |||
[[File:Strainseeker flowchart.png]] | |||
===Computational Phenotyping using CARD and VFDB Databases=== | |||
The Comprehensive Antibiotic Resistance Database (CARD) includes information on resistant genes, proteins coded by those genes, and their associated phenotypes. Since we want to understand the cause of heteroresistance and/or heterosusceptibility, we performed computational phenotyping against the CARD database to determine which antibiotic genes were present within the genome assembly. | |||
The Virulence Factors Database (VFDB) is a reference database that holds information on virulence factors of pathogenic bacteria. They hold about 2,353 virulence factors including bacterial toxins, cell surface proteins, cell surface carbohydrates, and hydrolytic enzymes that may contribute to the pathogenicity of the bacterium. Computational phenotyping was performed against the VFDB database as well. | |||
[[File:Card flowchart.png]] | |||
[[File:Vfdb.png]] | |||
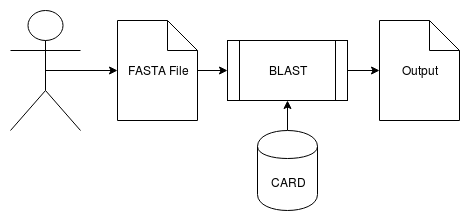
There are two pipelines for processing input FASTA file. For CARD, we use BLAST and filter its results based on % coverage and % identity to categorize antibiotic resistance genes as "High", "Medium", "Low". These labels represent our confidence in the BLAST results being antibiotic resistance genes. This output is parsed further to categorize each gene into drug and resistance mechanism category. The final outputs for the CARD pipeline are barcharts that shows the number of counts for antibiotics and resistance mechanisms for a given input FASTA file. | |||
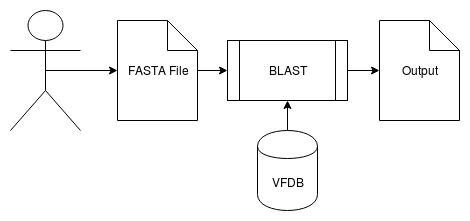
Similar to CARD, we use BLAST to retrieve information for a given FASTA file. This pipeline displays a table of virulence factor found in the genome. | |||
===pyani=== | |||
In order to calculate the Average Nucleotide Identity (ANI) between genomes, we implemented the python tool pyani. ANI is a measure of genome relatedness, and it shows how many nucleotides are identical between two genomes. The ANI value is related to DNA-DNA hybridization values, which traditionally indicate the microbial species definition. ANI values above 95% indicate that two genomes are the same species. | |||
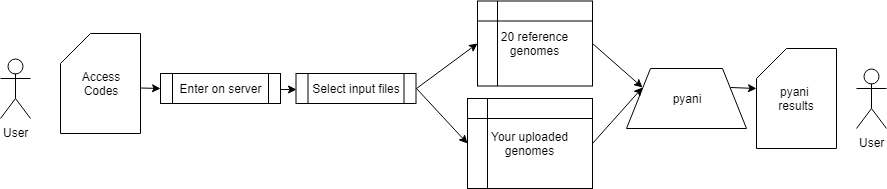
We implemented pyani through using a very quick alignment tool - mummer. In our server, we run pyani between six genomes. The user is able to choose among 20 reference genomes and any genome that a user has uploaded. So, ANI can be used to see similarities and differences between a dataset and also to get an idea of identity to ''Klebsiella'' references. The result from pyani is then parsed to generate a heatmap with the input genomes. | |||
[[File:Pyani_updated.png]] | |||
==Webserver ([http://predict2018a.biosci.gatech.edu/ Link])== | |||
===Upload File=== | |||
This is where the users can upload files to analyze, such as genome FASTA file. The users will be given an access code, which will be used later to retrieve their uploaded files for analysis available on the webserver. | |||
===Assembly=== | |||
Aside from performing assembly, under dropdown menu for Assembly, there is a download page from which the users can access their assembled files. Users will be sent an email and download code when the job is complete. | |||
===Predictions=== | |||
By clicking the dropdown menu, the prediction pipelines that were made available for the webserver is shown. Currently available ones include CARD/VFDB analysis, pyani, and StrainSeeker. | |||
===Screenshots=== | |||
[[File:Home team1.PNG]] | |||
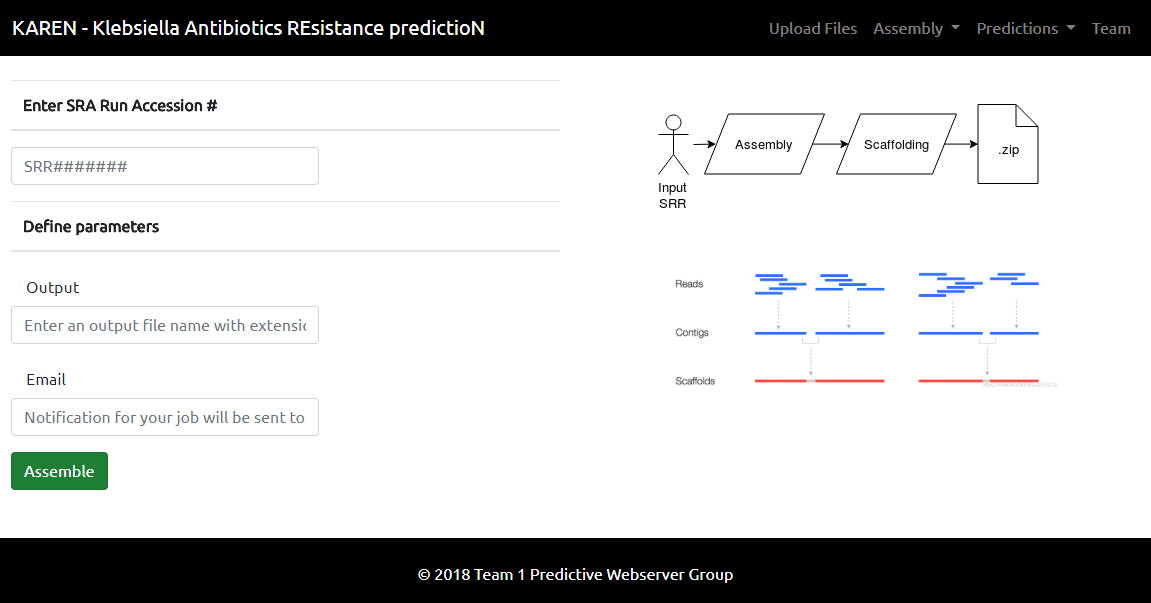
[[File:Assembly team1.PNG]] | |||
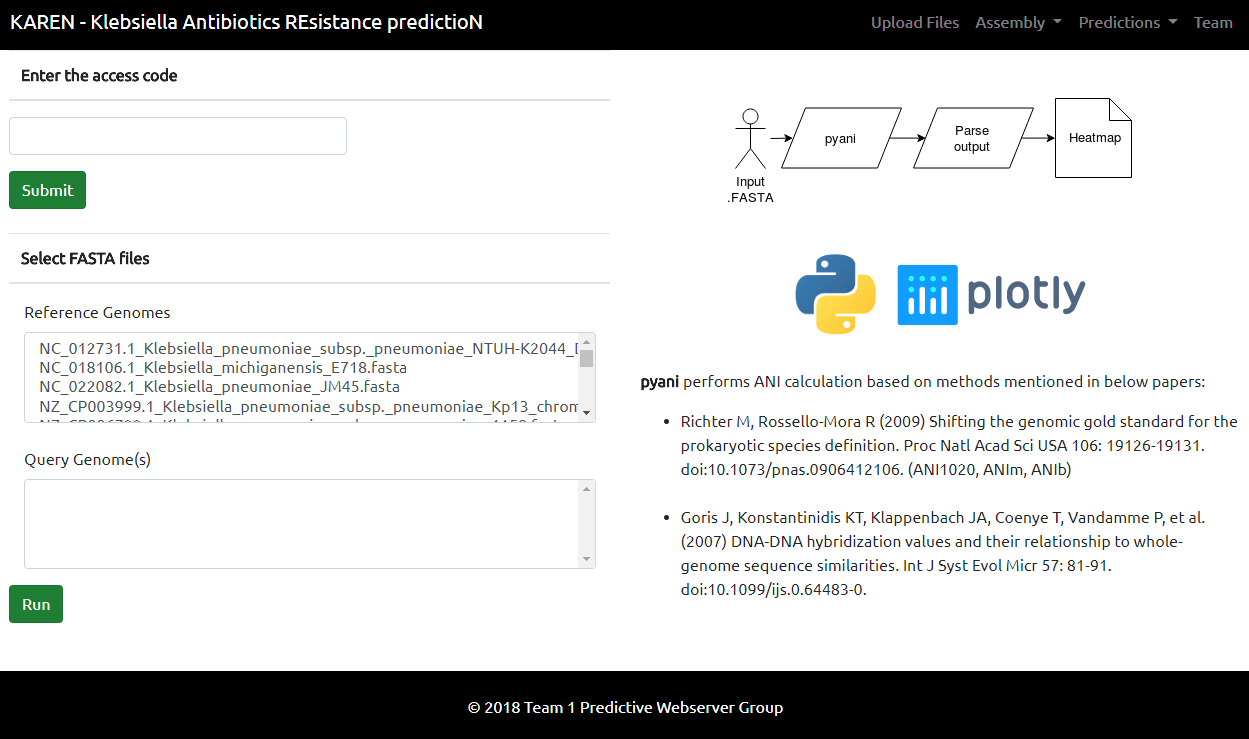
[[File:Pyani team1.PNG]] | |||
=References= | |||
Andrews, S. (2010). ''FastQC: a quality control tool for high throughput sequence data''. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc | |||
Bolger, A. M., Lohse, M., & Usadel, B. (2014). ''Trimmomatic: A flexible trimmer for Illumina Sequence Data''. Bioinformatics, btu170. | |||
Chen, L., Yang, J., Yu, J., Yao, Z., Sun, L., Shen, Y., Jin, Q. (2005). ''VFDB: a reference database for bacterial virulence factors'' .Nucleic Acids Res. 33:D325-8. | |||
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, et al. (2007). ''DNA-DNA hybridization values and their relationship to whole-genome sequence similarities''. Int J Syst Evol Micr 57: 81-91. doi:10.1099/ijs.0.64483-0. | |||
Jia et al. (2017). ''CARD 2017: expansion and model-centric curation of the Comprehensive Antibiotic Resistance Database''. Nucleic Acids Research, 45, D566-573 | |||
Leighton, Pritchard: The James Hutton Institute (2015). ''PyANI''. https://github.com/widdowquinn/pyani | |||
Roosaare et al. (2017). ''StrainSeeker: fast identification of bacterial strains from raw sequencing reads using user-provided guide trees''. PeerJ 5:e3353 | |||
Latest revision as of 11:31, 28 April 2018
Introduction
Background
The objectives of our BIOL 7210: Computational Genomics teams were to, given unassembled genome sequence data from the Weiss Lab at the Emory University School of Medicine, proceed through five distinct stages of analysis and interpretation of that data: genome assembly, gene prediction, functional annotation, comparative genomics, and production of a predictive webserver. At the last stage, our goal was to create a predictive webserver that performed the functionalities of some, if not all, of the work from previous groups.
Goals
Our goals for a predictive webserver were as follows:
- Assemble input reads
- Analyze assemblies
- Visualize results in user-friendly format
- Implement a way for results to be downloaded
KAREN
Klebsiella Antibiotics REsistance PredicitioN (KAREN) is a culmination of these objectives and is able to perform the following analyses given an input of raw sequence reads:
- De novo assembly
- Species identification
- Strain identification
- Average Nucleotide Identity
- Computational phenotyping
- Visualization of results
Technologies Used
For the creation and development of this webserver, we used PHP framework for server-side programming. PHP provides a strong frameworks to support MySQL and Apache Server. Also, PHP provides the feasibility of the development of Model-View-Controller (MVC) framework, which provides a more simple user-interface. There are many MVC frameworks available, among which we used Laravel. Laravel was used because of its extensive documentation and a large community of developers. It also currently is one of the most widely used PHP frameworks. For more information, please visit their website
This webserver is built on PHP v7.0.0 and Laravel v5.5.
Functionalities
De novo Genome Assembly using SKESA
We used SKESA for de novo genome assembly. The input to the assembler was raw reads (forward & reverse) retrieved using SRA accession numbers. The output contigs then were scaffolded using a tool called SSPACE. For more information on the assembly pipeline, refer to Genome Assembly team. SKESA is currently unpublished.
Species & Strain Typing by StrainSeeker
Strainseeker is a tool which lets you rapidly and accurately make an assessment of the species and strain of a bacterial assembly. StrainSeeker has a pre-built database that is uses for species identification and works on paired-end reads to identify strain type. It has the ability to identify novel strains and is therefore a useful tool for further assessment of a sample of unknown origin.
perl builder.pl -n refseq_guide_tree.nwk -d strain_fasta_directory -w 32 -o my_database
perl seeker.pl -i sample_file.fastq -d ss_db_w32 -o sample_result.txt
The pipeline works by taking a FASTA file as an input, which will then be processed by StrainSeeker. The output will be parsed by one of the scripts in the pipeline and visualization with tables containing information is generated.
Computational Phenotyping using CARD and VFDB Databases
The Comprehensive Antibiotic Resistance Database (CARD) includes information on resistant genes, proteins coded by those genes, and their associated phenotypes. Since we want to understand the cause of heteroresistance and/or heterosusceptibility, we performed computational phenotyping against the CARD database to determine which antibiotic genes were present within the genome assembly.
The Virulence Factors Database (VFDB) is a reference database that holds information on virulence factors of pathogenic bacteria. They hold about 2,353 virulence factors including bacterial toxins, cell surface proteins, cell surface carbohydrates, and hydrolytic enzymes that may contribute to the pathogenicity of the bacterium. Computational phenotyping was performed against the VFDB database as well.
There are two pipelines for processing input FASTA file. For CARD, we use BLAST and filter its results based on % coverage and % identity to categorize antibiotic resistance genes as "High", "Medium", "Low". These labels represent our confidence in the BLAST results being antibiotic resistance genes. This output is parsed further to categorize each gene into drug and resistance mechanism category. The final outputs for the CARD pipeline are barcharts that shows the number of counts for antibiotics and resistance mechanisms for a given input FASTA file.
Similar to CARD, we use BLAST to retrieve information for a given FASTA file. This pipeline displays a table of virulence factor found in the genome.
pyani
In order to calculate the Average Nucleotide Identity (ANI) between genomes, we implemented the python tool pyani. ANI is a measure of genome relatedness, and it shows how many nucleotides are identical between two genomes. The ANI value is related to DNA-DNA hybridization values, which traditionally indicate the microbial species definition. ANI values above 95% indicate that two genomes are the same species.
We implemented pyani through using a very quick alignment tool - mummer. In our server, we run pyani between six genomes. The user is able to choose among 20 reference genomes and any genome that a user has uploaded. So, ANI can be used to see similarities and differences between a dataset and also to get an idea of identity to Klebsiella references. The result from pyani is then parsed to generate a heatmap with the input genomes.
Webserver (Link)
Upload File
This is where the users can upload files to analyze, such as genome FASTA file. The users will be given an access code, which will be used later to retrieve their uploaded files for analysis available on the webserver.
Assembly
Aside from performing assembly, under dropdown menu for Assembly, there is a download page from which the users can access their assembled files. Users will be sent an email and download code when the job is complete.
Predictions
By clicking the dropdown menu, the prediction pipelines that were made available for the webserver is shown. Currently available ones include CARD/VFDB analysis, pyani, and StrainSeeker.
Screenshots
References
Andrews, S. (2010). FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Bolger, A. M., Lohse, M., & Usadel, B. (2014). Trimmomatic: A flexible trimmer for Illumina Sequence Data. Bioinformatics, btu170.
Chen, L., Yang, J., Yu, J., Yao, Z., Sun, L., Shen, Y., Jin, Q. (2005). VFDB: a reference database for bacterial virulence factors .Nucleic Acids Res. 33:D325-8.
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, et al. (2007). DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Micr 57: 81-91. doi:10.1099/ijs.0.64483-0.
Jia et al. (2017). CARD 2017: expansion and model-centric curation of the Comprehensive Antibiotic Resistance Database. Nucleic Acids Research, 45, D566-573
Leighton, Pritchard: The James Hutton Institute (2015). PyANI. https://github.com/widdowquinn/pyani
Roosaare et al. (2017). StrainSeeker: fast identification of bacterial strains from raw sequencing reads using user-provided guide trees. PeerJ 5:e3353