Team II Genome Assembly Group: Difference between revisions
| Line 42: | Line 42: | ||
5. Sequence length distribution | 5. Sequence length distribution | ||
We use ''FastQC'' for receiving these information and making some visualization. These kind of information will give us as overall view and help us to improve the quality of out sequences as raw inputs for the assemblers if it is necessary. For example, The presence of poor quality or technical sequences such as adapters in next-generation sequencing (NGS) data can easily the quality of assembly. We used ''Trim Galore'' and ''Sickle'' to guarantee the best performance for each assembly method that will use. | We use '''FastQC''' [5] for receiving these information and making some visualization. These kind of information will give us as overall view and help us to improve the quality of out sequences as raw inputs for the assemblers if it is necessary. For example, The presence of poor quality or technical sequences such as adapters in next-generation sequencing (NGS) data can easily the quality of assembly. We used ''Trim Galore'' and ''Sickle'' to guarantee the best performance for each assembly method that will use. | ||
'''Trim Galore!''' is a wrapper script to automate quality and adapter trimming as well as quality control, with some added functionality to remove biased methylation positions for RRBS sequence files [ | '''Trim Galore!''' is a wrapper script to automate quality and adapter trimming as well as quality control, with some added functionality to remove biased methylation positions for RRBS sequence files [6]. | ||
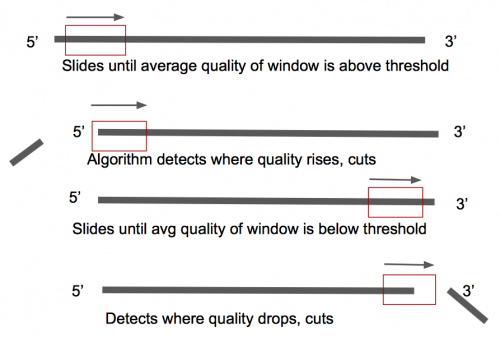
'''Sickle''' is a tool that uses sliding windows along with quality and length thresholds to determine when quality is sufficiently low to trim the 3'-end of reads and also determines when the quality is sufficiently high enough to trim the 5'-end of reads (Figure 3) [ | '''Sickle''' is a tool that uses sliding windows along with quality and length thresholds to determine when quality is sufficiently low to trim the 3'-end of reads and also determines when the quality is sufficiently high enough to trim the 5'-end of reads (Figure 3) [7]. | ||
[[File: 5.png | 500px]] | [[File: 5.png | 500px]] | ||
| Line 60: | Line 60: | ||
After trimming the adapter and the low quality reads, we can check the quality of all sequences at one with the ''MultiQC'' tool. | After trimming the adapter and the low quality reads, we can check the quality of all sequences at one with the ''MultiQC'' tool. | ||
'''MultiQC''' is a general use tool, perfect for summarising the output from numerous bioinformatics tools [ | '''MultiQC''' is a general use tool, perfect for summarising the output from numerous bioinformatics tools [8]. | ||
[[File: 5MULTIQC.png | 650px]] | [[File: 5MULTIQC.png | 650px]] | ||
Revision as of 14:34, 5 March 2018
Introduction
Background
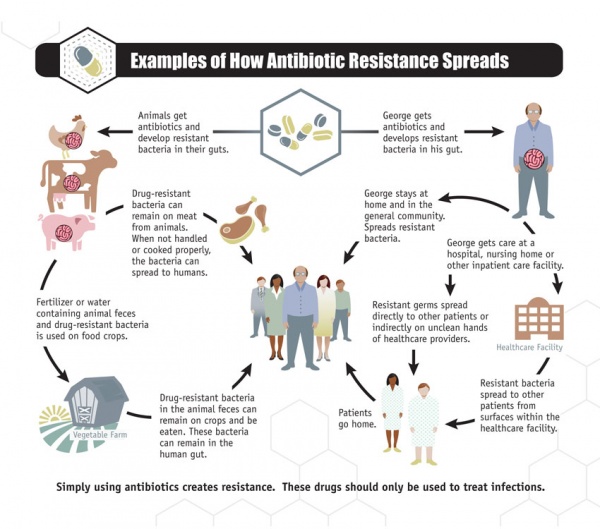
Antibiotic resistance has been called one of the world’s most pressing public health problems. Antibiotic resistance is the ability of bacteria to resist the effects of an antibiotic. It occurs when bacteria change in a way that reduces the effectiveness of drugs, chemicals, or other agents designed to cure or prevent infections. The bacteria survive and continue to multiply, causing more harm (Figure 1.a.). Antibiotic resistance can cause illnesses that were once easily treatable with antibiotics to become dangerous infections, prolonging suffering for children and adults. Antibiotic-resistant bacteria can spread to family members, schoolmates, and co-workers, and may threaten your community (Figure 1.b). Antibiotic-resistant bacteria are often more difficult to kill and more expensive to treat and in some cases, can lead to serious disability or even death [1].
Figure 1: a. How antibiotic resistance happens, b. How antibiotic resistance spreads
Data
Emory Antibiotic Resistance Center (ARC) in Emory University, School of medicine tries to better understand antibiotic resistance to combat this crisis and improve human health. Their goals include learning how antibiotic resistance develops, optimizing the way antibiotics are used to preserve their power, and discovering novel therapeutics and vaccines to directly combat antibiotic-resistant pathogens. Solving the crisis of antibiotic resistance requires a multi-faceted approach that crosses traditional boundaries [2].
They Provided us with a sample of 262 pair-end raw reads sequencing of Klebsiella spp, from illumina MiSeq.
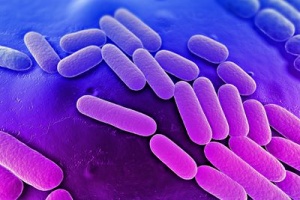
Klebsiella is a genus of nonmotile, Gram-negative, oxidase-negative, rod-shaped bacteria (Figure 2) with a prominent polysaccharide-based capsule. Klebsiella species are found everywhere in nature. The members of the genus Klebsiella are a part of the human and animal's normal flora in the nose, mouth and intestines. The species of Klebsiella are all gram-negative and non-motile. They tend to be shorter and thicker when compared to others in the Enterobacteriaceae family. The cells are rods in shape and generally measures 0.3 to 1.5 µm wide by 0.5 to 5.0 µm long. They can be found singly, in pairs, in chains or linked end to end. Klebsiella can grow on ordinary lab medium and do not have special growth requirements, like the other members of Enterobacteriaceae. Some of Klebsiella types are: K.granulomatis, K. oxytoca, K. michiganensis and K. pneumoniae (type-species: K. p. subsp. ozaenae, K. p. subsp. pneumoniae, K. p. subsp. rhinoscleromatis) [3]. Though, they have been extensively studied, for example, only four complete genomes of K. pneumoniae were available till 2011 [4]. To better understand the multidrug resistance factors in Klibsiella, we need to determine genome DNA sequences of strains.
Figure 2: Scanning electron microscope image of Klebsiella pneumoniae. From: Bioquell.com
Objectives
- To distinguish between susceptible and heteroresistant strains/species
- To discover genomic determinants of antibiotic resistance
- To develop a predictive web server
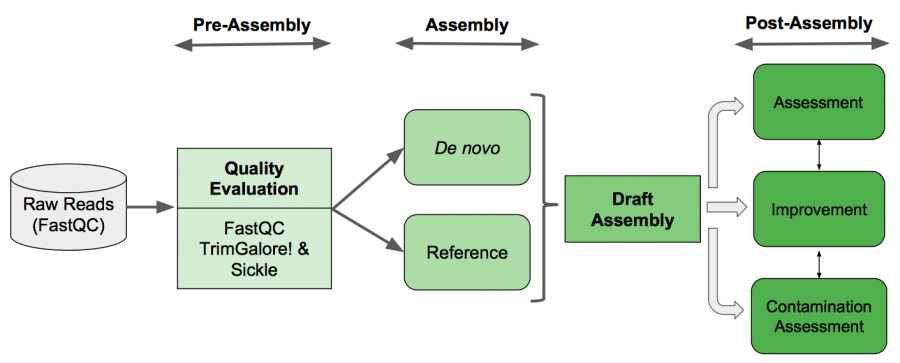
Genome Assembly Pipeline
Pre-Assembly
Before doing any assembly, we would like to got some basic statistics about our dataset as:
1. Per base sequence quality
2. Per base N content
3. Per base GC content
4. Overrepresented sequences
5. Sequence length distribution
We use FastQC [5] for receiving these information and making some visualization. These kind of information will give us as overall view and help us to improve the quality of out sequences as raw inputs for the assemblers if it is necessary. For example, The presence of poor quality or technical sequences such as adapters in next-generation sequencing (NGS) data can easily the quality of assembly. We used Trim Galore and Sickle to guarantee the best performance for each assembly method that will use.
Trim Galore! is a wrapper script to automate quality and adapter trimming as well as quality control, with some added functionality to remove biased methylation positions for RRBS sequence files [6].
Sickle is a tool that uses sliding windows along with quality and length thresholds to determine when quality is sufficiently low to trim the 3'-end of reads and also determines when the quality is sufficiently high enough to trim the 5'-end of reads (Figure 3) [7].
Figure 3: How Sickle is working
We used the following parameters for Trim Galore and Sickle and the results for before and after trimming shown in Figure 4.
Figure 4: Per base sequence quality and adapter content of one of the sample before and after trimming
After trimming the adapter and the low quality reads, we can check the quality of all sequences at one with the MultiQC tool.
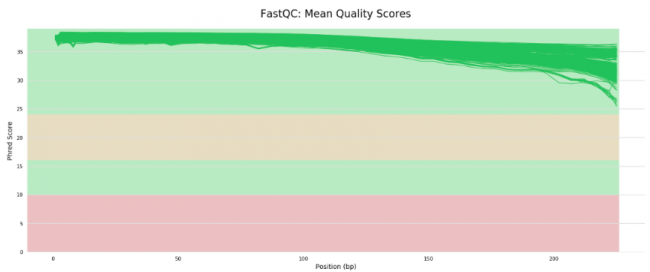
MultiQC is a general use tool, perfect for summarising the output from numerous bioinformatics tools [8].
Figure 5: Mean quality scores
Assembly
In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. This is needed as DNA sequencing technology cannot read whole genomes in one go, but rather reads small pieces of between 20 and 30000 bases, depending on the technology used [8].
Reference Assembly
A type of genome assembly where the reads are mapped (or compared) to a known version of the organism’s genome. It saves time and is significantly more accurate and assemble the genome with less contigs. But, it has some disadvantage too. We would not discover new genes with this method of assembly, new (completely different) sequences may lost. It requires a reference that is very similar to the sequenced data. If multiple positions on the reference genome are equally likely for a read, then, reads are ignored, placed at multiple locations or placed at the first likely position.
As we don't know the species of the Klebsiella, cannot simply choose a Klebsiella species to use as reference genome. Therefore, we test conservation between species with ANI and Mauve tools. Then, attempt to determine the species of each sample with StrainSeeker. At the end, assemble samples using test reference genomes with Bowtie and SMALT.
StrainSeeker is a program for detecting bacterial strains from raw sequencing reads. It gives us the option of fully customizable database of our own strains of interest. It can detect novel strains that are related to strains in the database and quickly handle large amounts of data.
Making database ...
So, for each single read we receive a list of references and we pick the closest one for each and categorize all the reads to different bin and then assemble that batch based on the assigned reference. We can illustrate the steps with the cartoon in Figure 5.
Figure 5: Strain Seeker
De Novo Assembly
De novo sequencing refers to sequencing a novel genome where there is no reference sequence available for alignment. Sequence reads are assembled as contigs, and the coverage quality of de novo sequence data depends on the size and continuity of the contigs (ie, the number of gaps in the data).