Team I Gene Prediction Group
Team members: Genevieve Brandt, Victoria Caban, Yuntian He, Junyu Li, Yiqiuyi Liu, Yihao Ou, Wenyi Qiu, Casey Smith, Mohit Thakur, Stephen Wist, Qinyu Yue
Introduction
Data
We were given de Novo assemblies of 258 isolates of Klebsiella spp..
Background
Our overarching goal is to understand what causes heteroresistance in Klebsiella spp. At this step, our objective was, given assembled genomes, to predict genes for Klebsiella that could later be annotated to understand functionality.
Gene Prediction
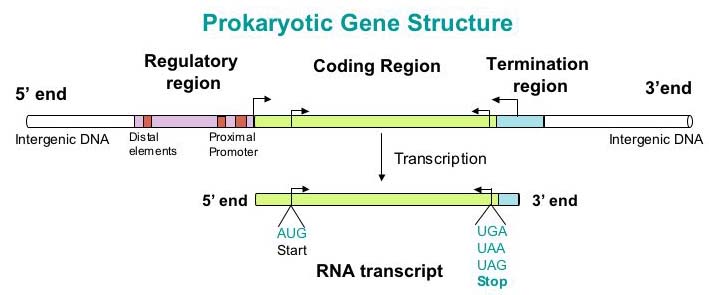
Gene prediction is the process of identifying the specific regions of genomic DNA that encode for genes. After sequencing and assembly, gene prediction is one of the first steps in understanding the genome of a species. In the past, confirming that the gene prediction is accurate demanded in vivo experimentation through gene knockout and other assays. Today, bioinformatics research has made it possible to predict the function of a gene based on its sequence alone. There are two general methods to do this: homology-based tools and ab-initio tools.
There is a big difference between prokaryotic and eukaryotic gene prediction. For eukaryotes, genes may be separated by introns, which makes it challenging to find the whole genomic sequence. Promoter sequences are more complex and less well understood in eukaryotes as well. In prokaryotes, the promoter regions are well understood, which is useful when using ab initio tools, since these tools search for signs of specific signs of protein coding genes. In prokaryotic genomes there are also contiguous open reading frames (ORFs) - when paired with the high amount of stop codons in prokaryotes this can indicate, with high probability, a gene being present. Our challenge when looking at ORFs is that every gene is a ORF, but not every ORF is a gene.
Methods
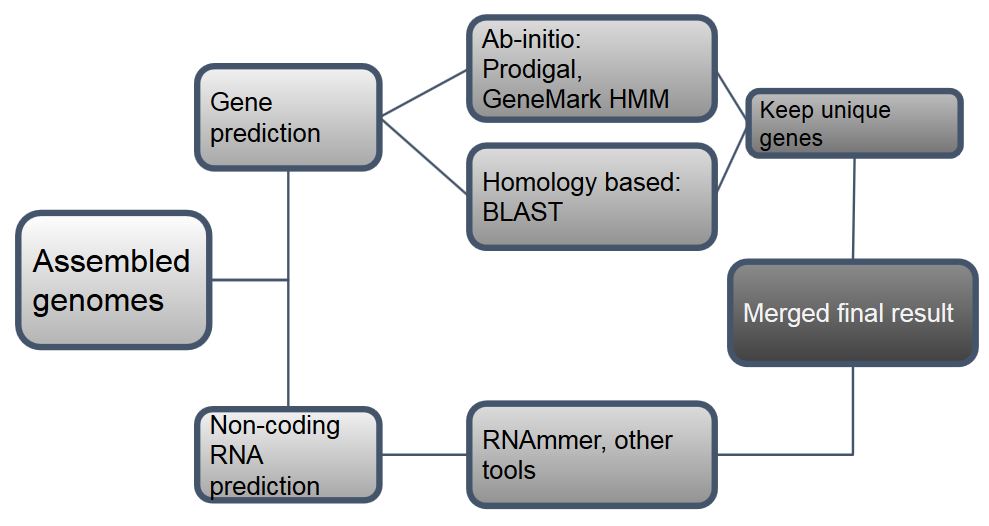
Pipeline (general workflow)
Homology-Based Methods
BLAST
One sequence by itself is not informative; it must be analyzed by comparative methods against existing sequence databases to develop hypothesis concerning relatives and function. BLAST(Basic Local Alignment Search Tool) is a widely used tool for similarity search, which adopts a Heuristic approach based on the Smith Waterman algorithm. BLAST provides not only the results of best local alignment, but various statistical significance as well.This algorithm compares a query sequence against reference sequence or specific database, such as nucleotide database. Because there are 258 assemblies that we are intending to BLAST, we chose to use local BLAST. The reference sequences we picked are the sequences which were used for gene assembly, the query sequences are the assembly results from Spades.
To run BLAST, we first need to create a database using the reference genome, then blast the query sequence against the database.
Commands:
makeblastdb -in reference.fasta -dbtype nucl -out db_name blastn -db db_name -query assembly_result.fasta -outfmt 6 -out blast_result.gff
Ab-initio Methods
Ab-initio tools always predict genes based on the gene content and signal detection(e.g. start/stop codon; Ribosome Biding Site, RBS etc.). In another words, they predict genes by analyzing statistical features of genes first, then separate the coding sequences and non-coding sequences apart.
There are so many different algorithms among different ab-initio tools. For example, Dynamic programming gene finding, Hidden Markov Model, Interpolated Markov Model, Linear Discriminant Analysis and so on. By comparing the sensitive, PPV and running time, we chose some tools below to generate our final results.
Prodigal
Pros:
Cons:
Parameters used:
Input: assembly file (.fna) Output: gene coordinates (.gff), nucleotide sequences (.fna), protein sequences (.fna) Command:
prodigal -i input.fna -o output.gff -f gff -d nucleotide_seq.fna -a protein_seq.fna
Reference: Hyatt, Doug, et al. "Prodigal: prokaryotic gene recognition and translation initiation site identification." BMC bioinformatics 11.1 (2010): 119.
Genemark.hmm
Before running GeneMark.hmm, it is necessary to got a mode file first. One way is to download the models from GeneMark website(http://opal.biology.gatech.edu/GeneMark/). Or you can generated them by GeneMarkS or GeneMark.
Pros:
Parameters used:
Input: assembly file (.fna) Output: gene coordinates (.gff), nucleotide sequences (.fna), protein sequences (.fna) Command:
gmhmmp -o <output.gff> -f G -m <model with its path> <input_file>
Reference: http://opal.biology.gatech.edu/GeneMark/background.html
Lukashin, Alexander V., and Mark Borodovsky. "GeneMark. hmm: new solutions for gene finding." Nucleic acids research 26.4 (1998): 1107-1115.
GLLIMMER
Run_glimmer.sh includes:
- Add GLIMMER executables to PATH
- Create output and log directories GFF/ and logs/
- GLIMMER calling, done through a pipeline script GLIMMER3 provides g3-from-scratch.csh
- Convert GLIMMER outputs to GFFs
Parameters used: (default in g3-from-scratch.csh)
Input: directory of assembly FASTA file(s) Output: Gene table (.predict and .detail) and byproducts (.icm, .train, .longorfs) Command:
bash run_glimmer.sh [directory]
Reference: https://ccb.jhu.edu/software/glimmer/glim302notes.pdf
ncRNA Methods
Infernal
Command:
cmscan --tblout <path to write tblout file> <cm file> <input fasta> > <path to write cmscan output>
Aragorn
Command:
./aragorn -t <input file>
RNAMMER
RNAmmer is a widely used ribosomal RNA prediction tool, it accepts both Prokaryotic and Eukaryotic inputs and returns result in gff, fasta and html formats based on need.
rnammer -S bac -m lsu,ssu,tsu -multi -gff output_file.gff -f output_file.fasta < input.fasta
Results
References
https://ghr.nlm.nih.gov/primer/basics/gene
https://www.biostat.wisc.edu/bmi776/spring-15/lectures/IMMs.pdf
http://ece.drexel.edu/gailr/ECE-S690-503/markov_models.ppt.pdf
http://onlinelibrary.wiley.com/doi/10.1042/BC20070137/full#footer-citing
https://iweb.langara.bc.ca/biology/mario/Biol2315notes/biol2315chap11.h